esiRNA coding human
The esiRNA technology is a powerful tool for loss of function gene studies. To illustrate the potency of silencing triggers quantitative real time (qRT)-PCR is often used to measure the knock-down rates. The knock-down validation on mRNA level was here performed 24 hours post transfection of esiRNA using qRT-PCR. To be able to assess knock-down rates, the expression levels of the mRNA of interest was compared to cells (HeLa or mouse ES) simultaneously transfected with Renilla Luciferase (negative control).
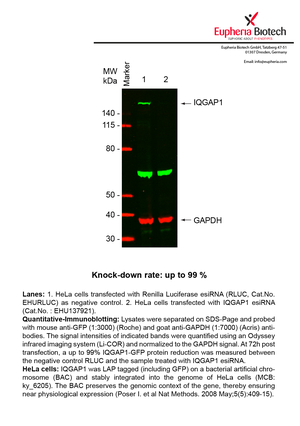
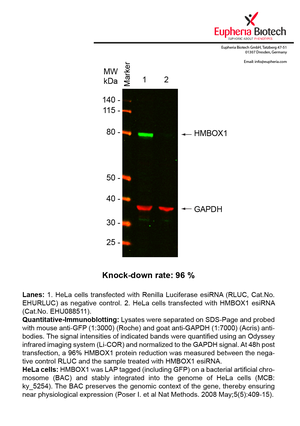
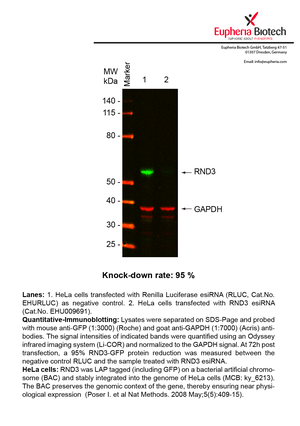
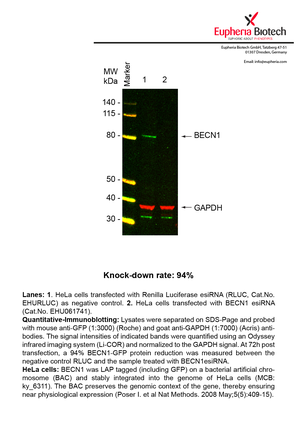
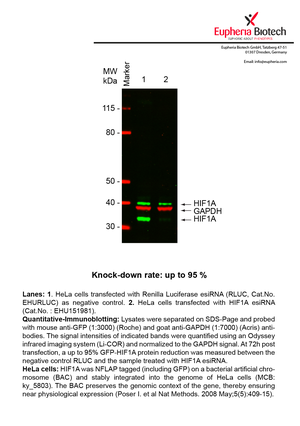
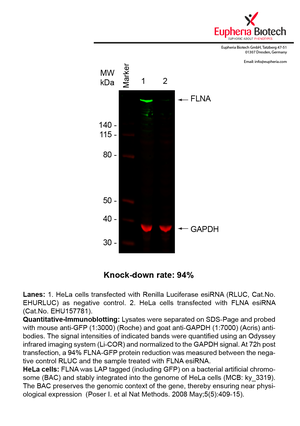
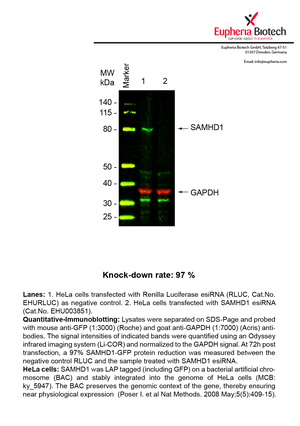
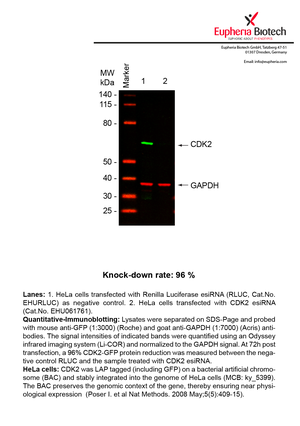
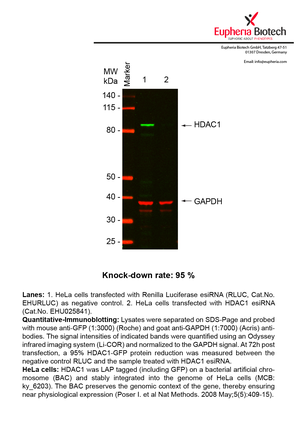
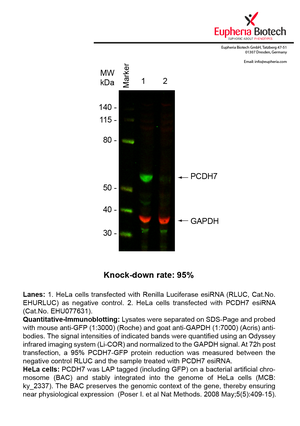
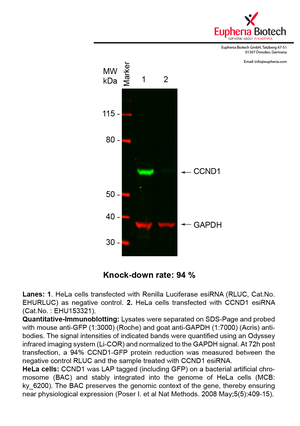
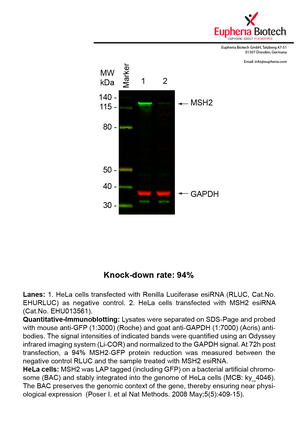
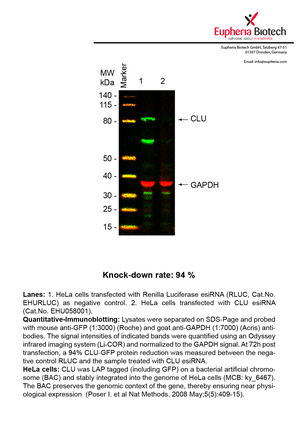
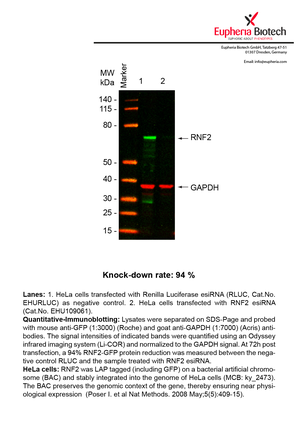
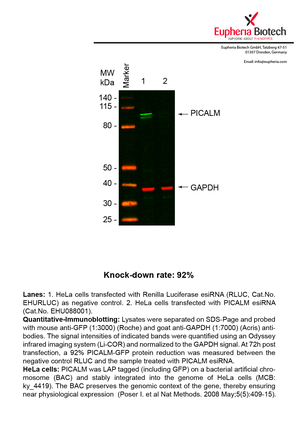
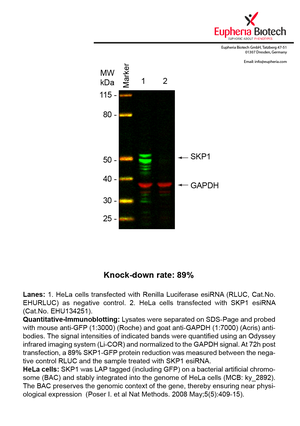
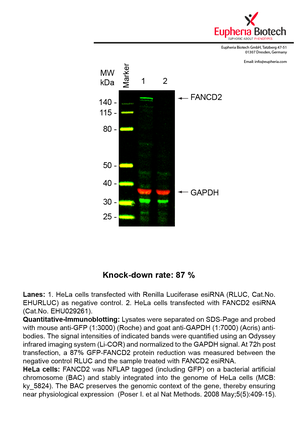
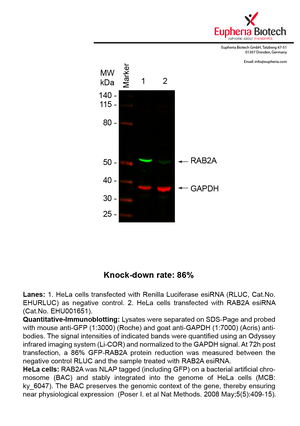
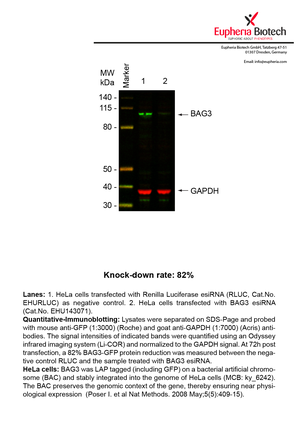
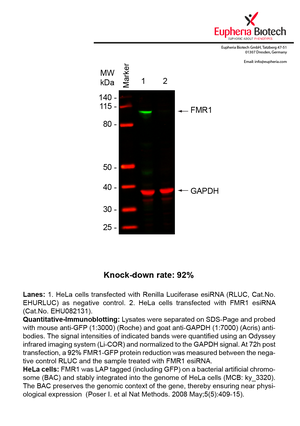
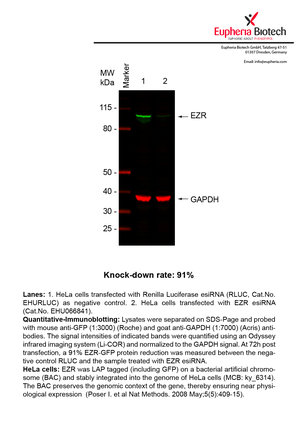
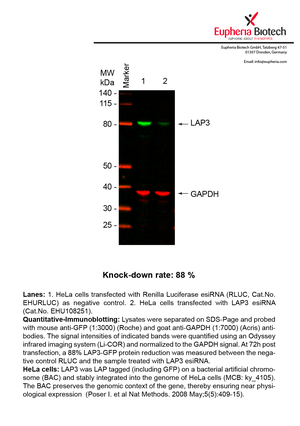
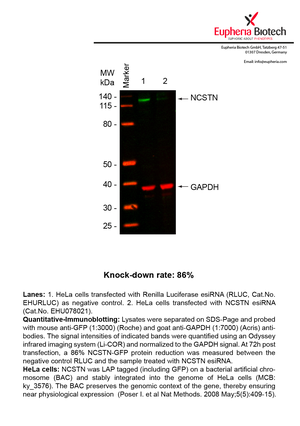
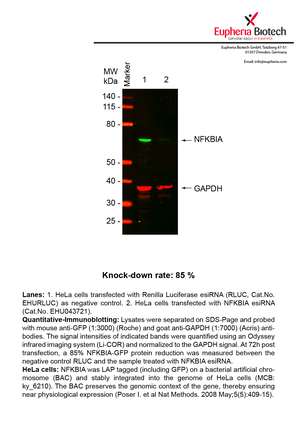
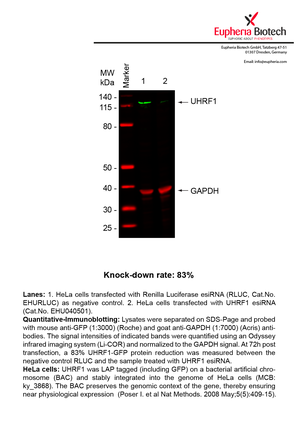
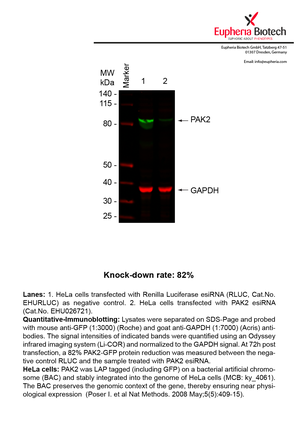
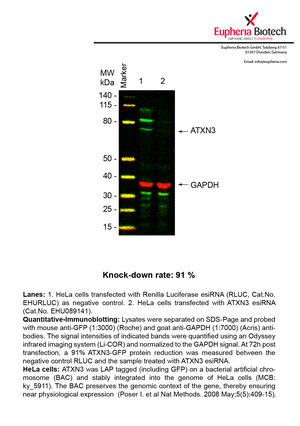
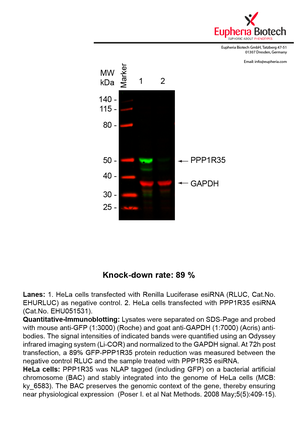
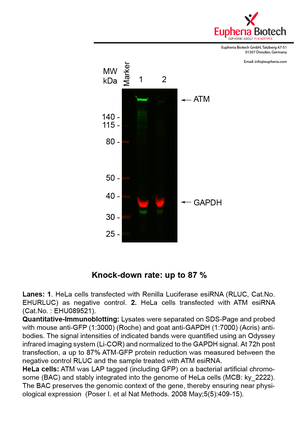
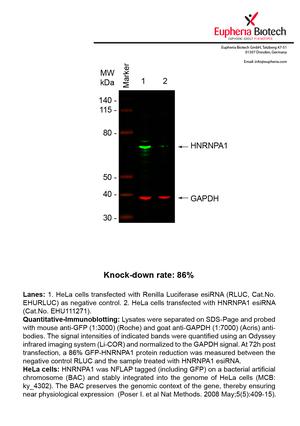
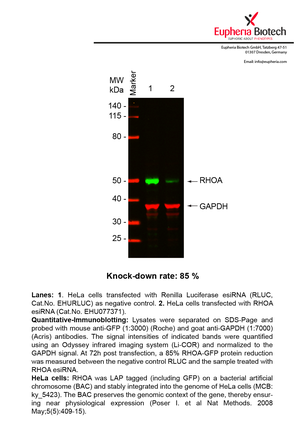
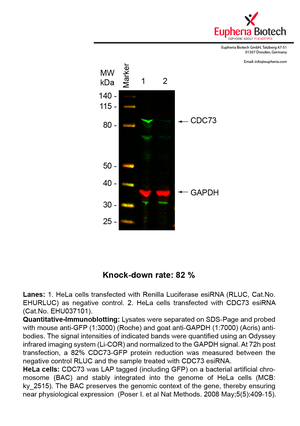
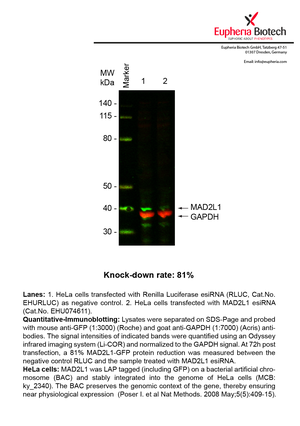
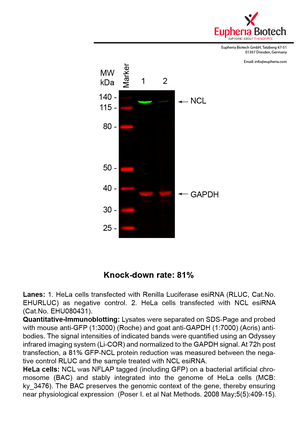
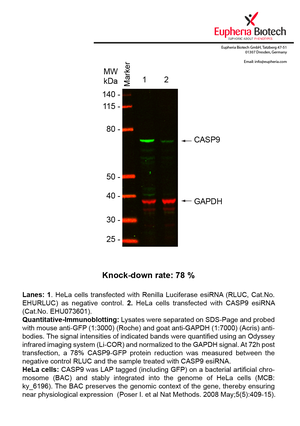
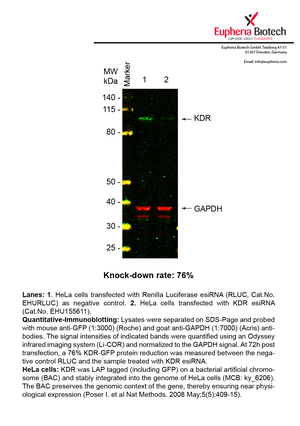
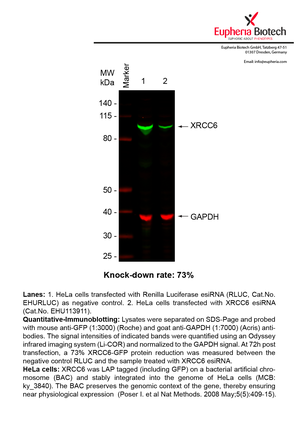
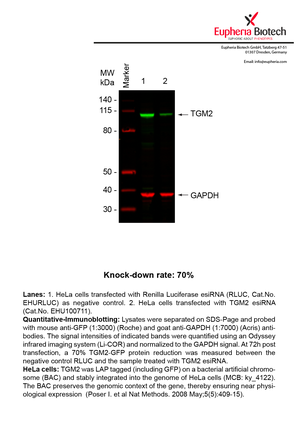
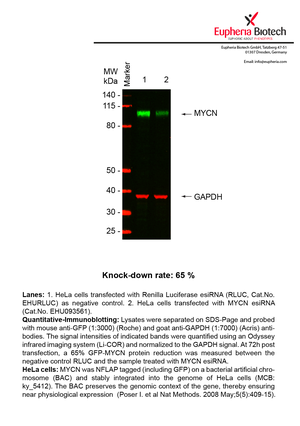
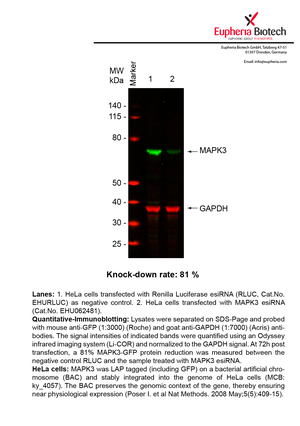
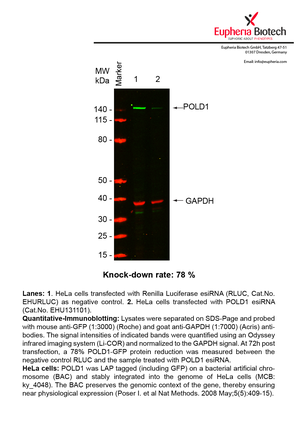
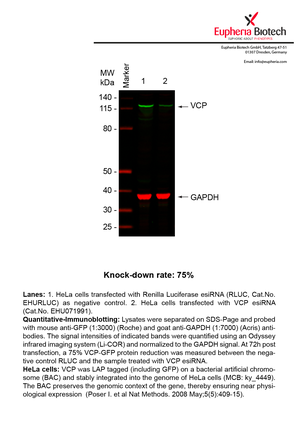
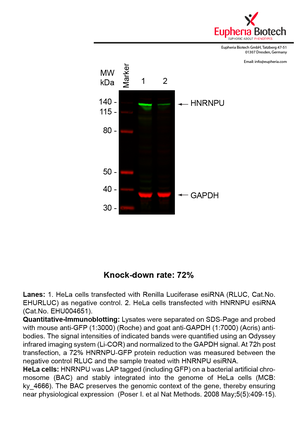
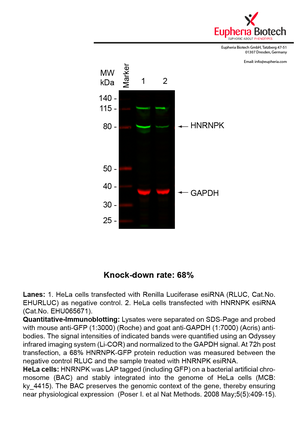
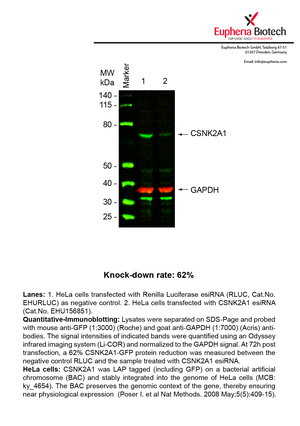
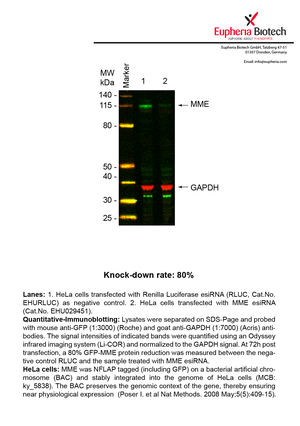
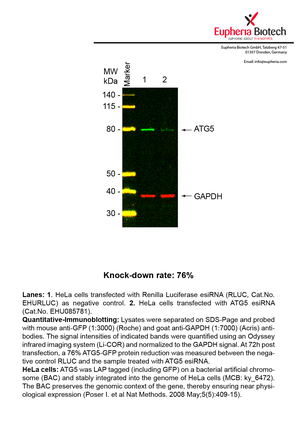
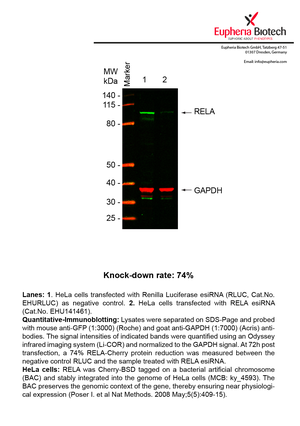
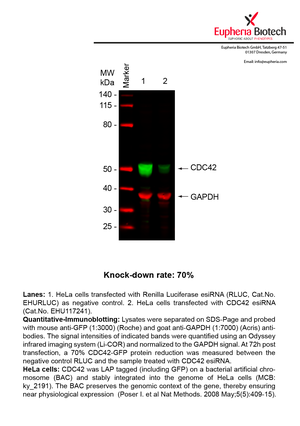
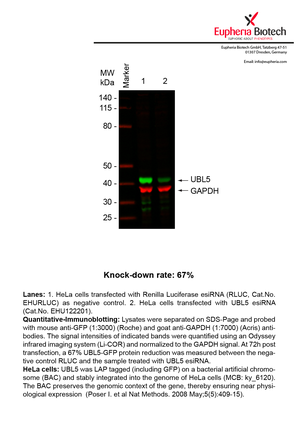
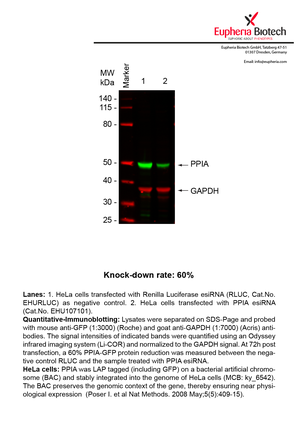
However, a more valuable measure of the knock-down potency in an RNAi experiment is the reduction in protein level. We therefore also validate the knock-down rates of esiRNAs on protein level using quantitative western blot analysis (Odyssey, Li-COR). The time point for maximum knock-down rate for each protein can vary significantly, as it is depending on factors such as protein-stability, turn-over rate or cell proliferation rates. The knock-down validation on protein level was here performed at 72 hours post transfection of esiRNA in HeLa cells. To be able to assess knock-down rates, the expression levels of the protein of interest was compared to HeLa cells simultaneously transfected with Renilla Luciferase (negative control).
Two approaches were used to monitor knock-down at protein level: 1. Specific antibodies for the protein of interest were used for the quantitative western blot analysis. 2. Proteins of interest were GFP-tagged on a bacterial artificial chromosome (BAC) and stably integrated into the genome of HeLa cells, allowing for near physiological expression (Poser I. et al Nat Methods. 2008 May;5(5):409-15). Using a GFP antibody the detection by quantitative western blot analysis of the protein of interest is straightforward.
| esiRNA ID | Gene Name | Gene Description | Ensembl ID | RefSeq ID | Knock-Down Rate | Pic / PDF | ||
| EHU137921 | IQGAP1 | IQ motif containing GTPase activating protein 1 | ENSG00000140575 | 99% | Pic / PDF | |||
| EHU003851 | SAMHD1 | SAM domain and HD domain 1 | ENSG00000101347 | NM_015474.3 | 97% | Pic / PDF | ||
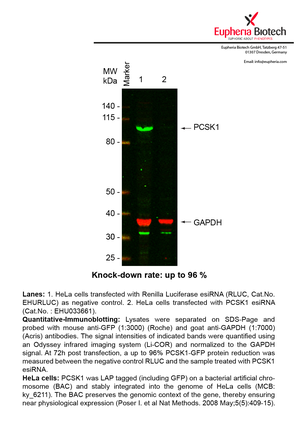
| EHU033661 | PCSK1 | proprotein convertase subtilisin/kexin type 1 | ENSG00000175426 | NM_000439 | 96% | Pic / PDF | ||
| EHU088511 | HMBOX1 | homeobox containing 1 | ENSG00000147421 | NM_024567 | 96% | Pic / PDF | ||
| EHU061761 | CDK2 | cyclin-dependent kinase 2 | ENSG00000123374 | NM_001798.3 NM_052827.2 | 96% | Pic / PDF | ||
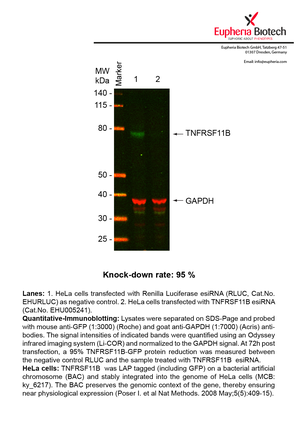
| EHU005241 | TNFRSF11B | tumor necrosis factor receptor superfamily, member 11b | ENSG00000164761 | NM_002546 | 95% | Pic / PDF | ||
| EHU009691 | RND3 | Rho family GTPase 3 | ENSG00000115963 | NM_005168 | 95% | Pic / PDF | ||
| EHU025841 | HDAC1 | histone deacetylase 1 | ENSG00000116478 | NM_004964 | 95% | Pic / PDF | ||
| EHU077631 | PCDH7 | protocadherin 7 | ENSG00000169851 | NM_032456.2 NM_001173523.1 NM_032457.3 NM_002589.2 | 95% | Pic / PDF | ||
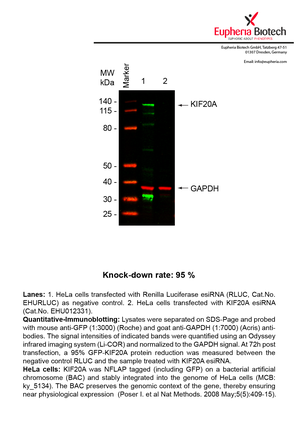
| EHU012331 | KIF20A | kinesin family member 20A | ENSG00000112984 | NM_005733.2 | 95% | Pic / PDF | ||
| EHU151981 | HIF1A | hypoxia inducible factor 1, alpha subunit (basic helix-loop-helix transcription factor) | ENSG00000100644 | 95% | Pic / PDF | |||
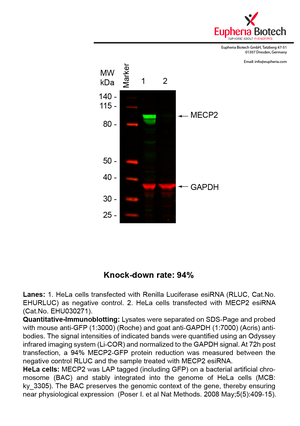
| EHU030271 | MECP2 | methyl CpG binding protein 2 (Rett syndrome) | ENSG00000169057 | NM_004992.3 | 94% | |||
| EHU061741 | BECN1 | beclin 1, autophagy related | ENSG00000126581 | NM_003766 | 94% | Pic / PDF | ||
| EHU153321 | CCND1 | cyclin D1 | ENSG00000110092 | NM_053056 | 94% | Pic / PDF | ||
| EHU058001 | CLU | clusterin | ENSG00000120885 | NR_045494.1 NM_001831.3 NR_038335.1 | 94% | Pic / PDF | ||
| EHU157781 | FLNA | filamin A, alpha | ENSG00000196924 | NM_001110556.1 NM_001456.3 | 94% | Pic / PDF | ||
| EHU013561 | MSH2 | mutS homolog 2, colon cancer, nonpolyposis type 1 (E. coli) | ENSG00000095002 | NM_001258281.1 NM_000251.2 | 94% | Pic / PDF | ||
| EHU109061 | RNF2 | ring finger protein 2 | ENSG00000121481 | NM_007212.3 | 94% | Pic / PDF | ||
| EHU088001 | PICALM | phosphatidylinositol binding clathrin assembly protein | ENSG00000073921 | NM_001206946.1 NM_007166.3 NM_001008660.2 NM_001206947.1 | 92% | Pic / PDF | ||
| EHU082131 | FMR1 | fragile X mental retardation 1 | ENSG00000102081 | NM_001185082.1 NM_001185081.1 NR_033700.1 NM_001185076.1 NM_001185075.1 NR_033699.1 NM_002024.5 | 92% | |||
| EHU089141 | ATXN3 | ataxin 3 | ENSG00000066427 | NM_030660 | 91% | |||
| EHU040791 | GSK3A | glycogen synthase kinase 3 alpha | ENSG00000105723 | NM_019884 | 91% | Pic / PDF | ||
| EHU066841 | EZR | ezrin | ENSG00000092820 | NM_001111077.1 NM_003379.4 | 91% | Pic / PDF | ||
| EHU051531 | PPP1R35 | protein phosphatase 1, regulatory subunit 35 | ENSG00000160813 | NM_145030.2 | 89% | Pic / PDF | ||
| EHU134251 | SKP1 | S-phase kinase-associated protein 1 | ENSG00000113558 | NM_170679.2 NM_006930.3 | 89% | |||
| EHU108251 | LAP3 | leucine aminopeptidase 3 | ENSG00000002549 | NM_015907.2 | 88% | |||
| EHU089521 | ATM | ataxia telangiectasia mutated | ENSG00000149311 | NM_000051 NM_138292 | 87% | Pic / PDF | ||
| EHU029261 | FANCD2 | Fanconi anemia, complementation group D2 | ENSG00000144554 | NM_001018115.1 NM_033084.3 | 87% | Pic / PDF | ||
| EHU078021 | NCSTN | nicastrin | ENSG00000162736 | NM_015331.2 | 87% | |||
| EHU111271 | HNRNPA1 | heterogeneous nuclear ribonucleoprotein A1 | ENSG00000135486 | NM_031157.2 | 86% | Pic / PDF | ||
| EHU001651 | RAB2A | RAB2A, member RAS oncogene family | ENSG00000104388 | NM_002865.2 | 86% | Pic / PDF | ||
| EHU043721 | NFKBIA | nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha | ENSG00000100906 | NM_020529 | 85% | Pic / PDF | ||
| EHU077371 | RHOA | ras homolog family member A | ENSG00000067560 | NM_001664 | 85% | Pic / PDF | ||
| EHU030431 | SNAI1 | snail homolog 1 (Drosophila) | ENSG00000124216 | NM_005985 | 83% | Pic / PDF | ||
| EHU040501 | UHRF1 | ubiquitin-like with PHD and ring finger domains 1 | ENSG00000034063 | NM_001048201.1 NM_013282.3 | 83% | |||
| EHU037101 | CDC73 | cell division cycle 73, Paf1 | ENSG00000134371 | NM_024529.4 | 82% | |||
| EHU143071 | BAG3 | BCL2-associated athanogene 3 | ENSG00000151929 | NM_004281.3 | 82% | Pic / PDF | ||
| EHU026721 | PAK2 | p21 protein (Cdc42/Rac)-activated kinase 2 | ENSG00000180370 | NR_027053.1 NM_002577.4 | 82% | Pic / PDF | ||
| EHU074611 | MAD2L1 | MAD2 mitotic arrest deficient-like 1 (yeast) | ENSG00000164109 | NM_002358.3 | 81% | Pic / PDF | ||
| EHU080431 | NCL | nucleolin | ENSG00000115053 | NM_005381.2 | 81% | |||
| EHU062481 | MAPK3 | mitogen-activated protein kinase 3 | ENSG00000102882 | NM_001040056 NM_002746 NM_001109891 | 81% | Pic / PDF | ||
| EHU029451 | MME | membrane metallo-endopeptidase | ENSG00000196549 | NM_007289.2 NM_007288.2 NM_000902.3 NM_007287.2 | 80% | Pic / PDF | ||
| EHU073601 | CASP9 | caspase 9, apoptosis-related cysteine peptidase | ENSG00000132906 | NM_001229,NM_032996 | 78% | Pic / PDF | ||
| EHU131101 | POLD1 | polymerase (DNA directed), delta 1, catalytic subunit 125kDa | ENSG00000062822 | 78% | Pic / PDF | |||
| EHU085781 | ATG5 | ATG5 autophagy related 5 | ENSG00000057663 | NM_004849 | 76% | Pic / PDF | ||
| EHU155611 | KDR | kinase insert domain receptor (a type III receptor tyrosine kinase) | ENSG00000128052 | NM_002253 | 76% | Pic / PDF | ||
| EHU071991 | VCP | valosin containing protein | ENSG00000165280 | NM_007126.3 | 75% | Pic / PDF | ||
| EHU141461 | RELA | v-rel reticuloendotheliosis viral oncogene homolog A (avian) | ENSG00000173039 | 74% | Pic / PDF | |||
| EHU113911 | XRCC6 | X-ray repair complementing defective repair in Chinese hamster cells 6 | ENSG00000196419 | NM_001469.3 | 73% | Pic / PDF | ||
| EHU004651 | HNRNPU | heterogeneous nuclear ribonucleoprotein U (scaffold attachment factor A) | ENSG00000153187 | NM_004501.3 NM_031844.2 | 72% | Pic / PDF | ||
| EHU117241 | CDC42 | cell division cycle 42 (GTP binding protein, 25kDa) | ENSG00000070831 | NM_044472.2 | 70% | |||
| EHU100711 | TGM2 | transglutaminase 2 (C polypeptide, protein-glutamine-gamma-glutamyltransferase) | ENSG00000198959 | NM_004613.2 | 70% | Pic / PDF | ||
| EHU065671 | HNRNPK | heterogeneous nuclear ribonucleoprotein K | ENSG00000165119 | NM_002140.3 NM_031262.2 NM_031263.2 | 68% | Pic / PDF | ||
| EHU122201 | UBL5 | ubiquitin-like 5 | ENSG00000198258 | NM_024292.3 | 67% | Pic / PDF | ||
| EHU093561 | MYCN | v-myc myelocytomatosis viral related oncogene, neuroblastoma derived (avian) | ENSG00000134323 | NM_005378 | 65% | Pic / PDF | ||
| EHU156851 | CSNK2A1 | casein kinase 2, alpha 1 polypeptide | ENSG00000101266 | NM_177560.2 NM_001895.3 NM_177559.2 NM_001256686.1 | 62% | Pic / PDF | ||
| EHU107101 | PPIA | peptidylprolyl isomerase A (cyclophilin A) | ENSG00000196262 | NM_021130.3 | 60% |